


Retired, Professor Emeritus.
No office at Hopkins
Email: [email protected]
B.S., Tufts University
Ph.D., University of California, Berkeley
Postdoctoral, Harvard University
Research Interests
As of 2019 I am no longer accepting new students.
Research in my laboratory was directed at understanding the basic biochemical and biophysical principles involved in protein function through the combined use of biochemistry, genetics, genetic engineering, and biophysics. Our criterion for understanding is that we can design and build systems that actually work and make use of these principles. Since we had extensive experience with the arabinose operon and systems related to it and developed a large collection of mutations in AraC and the regulatory region as well as many mutant DNA's and proteins most of our work was concentrated on this system. The ara system permits economic and rapid handling of the biology while displaying most of the repertoire of protein-protein, protein-DNA and gene regulatory principles that are found in prokaryotes and eukaryotes.
In 1984 we made the original discovery of DNA looping, a mechanism now known to be widely used in biology. Later we discovered the two domain structure to AraC and grew the crystals from which the structure of the dimerization domain in the presence and absence of arabinose was determined. This work in connection with biochemical and genetic studies led to the discovery of the role of the N-terminal arms on AraC and the mechanism where the two positions of the arms of the protein regulate the looping-unlooping activity of the protein. We demonstrated that a version of the mechanism can be ported to other proteins, and we constructed a β-galactosidase whose activity was controlled by the mechanism from AraC. The enzyme's activity is modulated by the presence of arabinose.
Wefound that the DNA binding domain of AraC may readily be overproduced and purified. It was a very good material for NMR studies, and we determined its structure by NMR. One recent objective was to determine precisely which residues are in contact between the dimerization and DNA binding domains both in the absence of arabinose and in the presence of arabinose. Genetic, and biochemical, methods were used to determine this. We found that it is the helicity of the interdomain linker that is controlled by the presence or absence of arabinose bound to AraC. In the absence of arabinose, the linker is helical, and in the presence of arabinose, it is a random coil.
Approaches that were commonly used in the laboratory include biochemistry, genetics, genetic engineering, physiological measurement, and biochemical and physical-chemical approaches, for example crystallography, fluorescence, electrophoresis, plasmon resonance, NMR, as well as computational approaches. Our primary, but not only, subject for comparison of theory and experiment was AraC protein.
Frequently we developed new experimental techniques to facilitate our studies. In the past we developed the DNA migration retardation assay so that biochemically meaningful information could be obtained from it and developed the missing contact method for determining specific amino acid-base interactions in DNA. More recently we developed methods for: locating linker regions in multi-domain proteins, constructing functional chimeric proteins when the domain locations are unknown, precise comparison of DNA binding affinities, and refolding DNA-binding proteins from insoluble inclusion bodies. We also developed a method for investigation of the very weak protein-protein and domain-domain interactions that are often found in complex regulatory systems.
Summary of the Regulation Mechanism of the Arabinose Operon
The gene products of the arabinose operon in Escherichia coli enable the cells to take up and catabolize the five carbon sugar, L-arabinose. In the absence of arabinose, the dimeric AraC protein actively represses its own synthesis and the synthesis of the AraB, AraA, and AraD gene
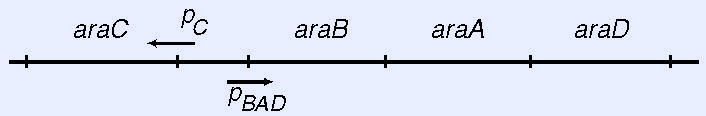
products by binding to the araO2 and araI1 half-sites and forming a DNA loop that blocks access of RNA polymerase to the pC and pBAD promoters. Upon the addition of arabinose, AraC ceases looping and binds instead to the adjacent half-sites, araI1 and araI2, where it and the cyclic AMP binding protein, CAP, both help RNA polymerase to bind to the pBAD promoter and speed the formation of open complex by RNA polymerase, thereby stimulating the synthesis of the AraB, AraA, and AraD gene products 100- to 500-fold.
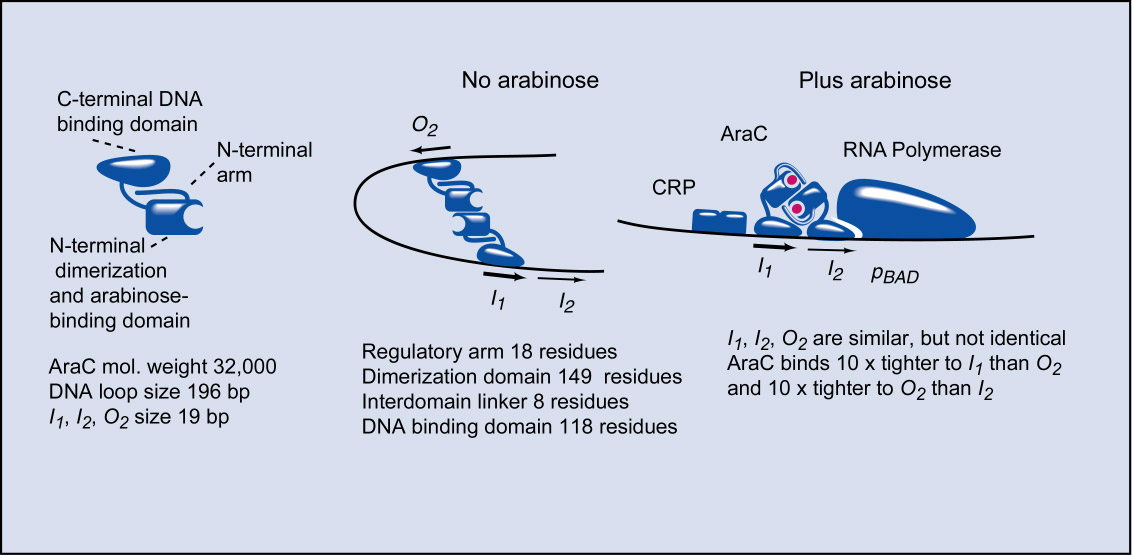
AraC protein consists of two connected domains, a DNA-binding domain that both binds to the various I-like sites and which also interacts with RNA polymerase to activate transcription, and a dimerization domain that also binds arabinose. AraC protein is primarily induced to form the DNA loop between the I1 and O2 half-sites by the N-terminal arms that extend from the dimerization domains. Precisely how they do this is unknown. For quite some time we thought it likely that the arms directly bound to the DNA binding domains and that this held the domains in orientations suitable for DNA looping. Multiple experiments cast doubt on this model and we now believe the arm's control of the DNA binding domains is an indirect influence on the rigidity of the interdomain linkers connecting the dimerization and DNA binding domains. Upon the binding of arabinose to the dimerization domains, the N-terminal arms reposition over the bound arabinose, ultimately resulting in allowing the interdomain linker to become a random coil, thus the DNA binding domains are released and are freed to assume any relative orientation they like. As a result, they now prefer to bind to the two adjacent, half-sites I1 and I2, where such binding activates transcription from pBAD.
Books and Recent Publications
Books
Practical Methods in Molecular Biology, Robert Schleif, Pieter Wensink, 1981, Springer-Verlag, New York. A cookbook of practical knowledge and methods used in molecular biology.
"Genetics and Molecular Biology" Johns Hopkins Press 2nd Ed. (1993). View or download the entire book in pdf format. (698 pages, 5.6 MB, Bookmarked). A graduate level textbook providing a rigorous and thoughtful presentation of the fundamentals of molecular biology.
"Analysis of Protein Structure and Function: A Beginner's Guide to CHARMM" Robert Schleif. View or download the entire book in pdf format (172 pages, 800 KB, Bookmarked). Describes the operation and use of CHARMM for molecular mechanics and molecular dynamics analysis of protein coordinates, energetics, and motions.
Scripts from "A Beginner's Guide to CHARMM" for downloading
"Elements of Biochemistry and Molecular Biology" Robert Schleif. This 100 page book in pdf format is written for advanced undergraduates or first year graduate students in Biology, Biochemistry, Chemistry, Physics, and Biophysics who desire a deeper understanding of some of the physical laws and principles that underlie much of Biology. The objective of the material is to help students develop a deeper understanding and an intuitive understanding of the principles involved. Ideal preparation for appreciating what the book offers would be completion of college level introductory courses in Calculus, Physics, Chemistry, and either Biology or Biochemistry. The book uses no mathematics beyond elementary calculus. Students’ appreciation of and understanding of some of the principles and phenomena will be assisted by their simulating phenomena on spreadsheets as developed and described in the text. These examples are described in sufficient detail that the spreadsheets may be constructed and utilized by novices. Throughout, interconnections between topics discussed in the book are noted as are connections to material not covered by the book. Approaches and strategies used in scientific research are both mentioned in passing and are inherent in the construction of much of the presentation.
Two Reviews of the Arabinose System
The AraC Protein: a Love-hate Relationship, Robert Schleif, BioEssays 25, 274-282 (2003)
Other Recent Papers
A Career's Work,the L-arabinose Operon: How it Functions and How We Learned it. EcoSal Plus 9:eESP-0012-(2021). https://doi.org/10.1128/ecosalplus.ESP-0012-2021.
Lab and Teaching
Laboratory Members
Graduate Students: All have finished
Laboratory Methods
Methods Cards These describe in recipe format how to do many routine molecular biology and biochemistry procedures. View or download the set of 160 pages that print on 5" x 8" index cards. Ours were kept in a recipe box and each category was printed on card stock of a different color.
Teaching
A Biophysical View of Biology, 2021.
Quantitative Biology, 2019, Four Lectures on Nucleic Acid and Protein--Structure and Function
Schleif's Portion of Graduate Biophysical Chemistry, 020.674.644, 2018
Advanced Molecular Biology, which became Foundations and Applications of Mol. Bio. (2017)
Some Comments for Graduate Students
Photographic: Some of my Better Pictures and A Few Tutorials
Best from 2011 and 2012 (10 images)
Venice and the Dolomites (12 images)
Scottish Highlands and Inner Hebrides (17 images)
From Our Bedroom Window (10 images)
Scenes from Around Hopkins (8 images)
From Fall to Winter in Maryland (5 images)
Longwood Botanic Gardens (8 images)
Iceland, primarily the Northern Coast (20 images)
Evenings around Baltimore, Towson, Fells Point, Hampden, Inner Harbor (12 images), 2018
Miscellaneous images, 2018 (9 images), 2018
Some computer art, 2019 (7 images)
Black and White 2020 (19 images)
Year of Covid Stay-at-home Images (21 images)
2021 A More Stay-at-home Images (15 images)
2021 B Still More Stay-at-home Images (15 images)
The Principles Behind Digital Image Sharpening
Gamma, Perception, Posterization, and Raw Conversion
Sensing Violet: The Human Eye and Digital Cameras
Random, Tennis and Science
Tennis Outcomes Related to Probability of Winning Individual Points
Optimum Serving and Receiving Strategies in Tennis and Competetive Games in General
Optimum Poaching Strategy in Doubles Tennis
Affinity of Transcription Factor-RNAP Interactions
A Very Simple Derivation of the Boltzmann Distribution
Ligand Binding to an Homo-oligomeric Protein, Cooperativity, Macro and Micro Dissociation Constants
Dimer Binding Affinity in Terms of Monomer Binding Affinity
How to Give Powerpoint Presentations from an iPhone
Importing Each Character of a Text String Into a Separate Cell of Excel